Research
Analysis of sexual reproduction in Closterium
Sexual reproduction is an essential process for almost all panmictic populations of organisms. Although organisms have different methods of sexual reproduction, there is a common basic process, i.e., two specialized sexually competent cells, which are determined genetically or developmentally, recognize each other and this leads to fertilization or conjugation.
1)Sexual reproductive processes in the heterothallic Closterium peracerosum-strigosum-littorale complex
The Closterium peracerosum-strigosum-littorale (C. psl.) complex is the best-characterized Closterium species. When mt+ (NIES-67) and mt– (NIES-68) cells (obtained from the National Institute for Environmental Studies, Ibaraki, Japan) are mixed in nitrogen-depleted medium in the presence of light, they divide once to form sexually competent gametangial cells. This cell division is called sexual cell division (SCD; 2). The gametangial cell of one mating type forms a pair with a cell of the other type (3). The paired cells then release their gametic protoplasts, which fuse immediately to form a zygospore (4-5).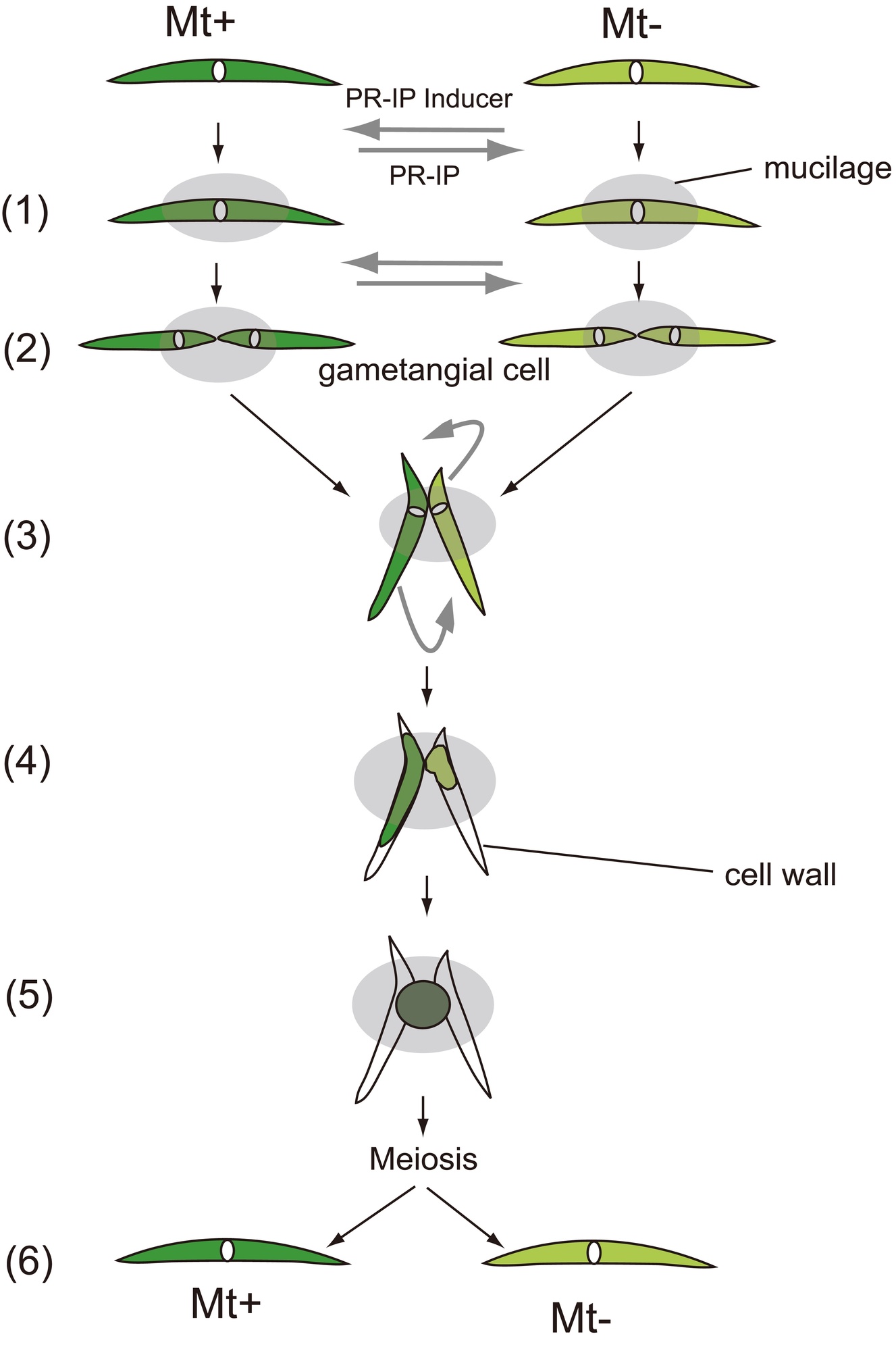
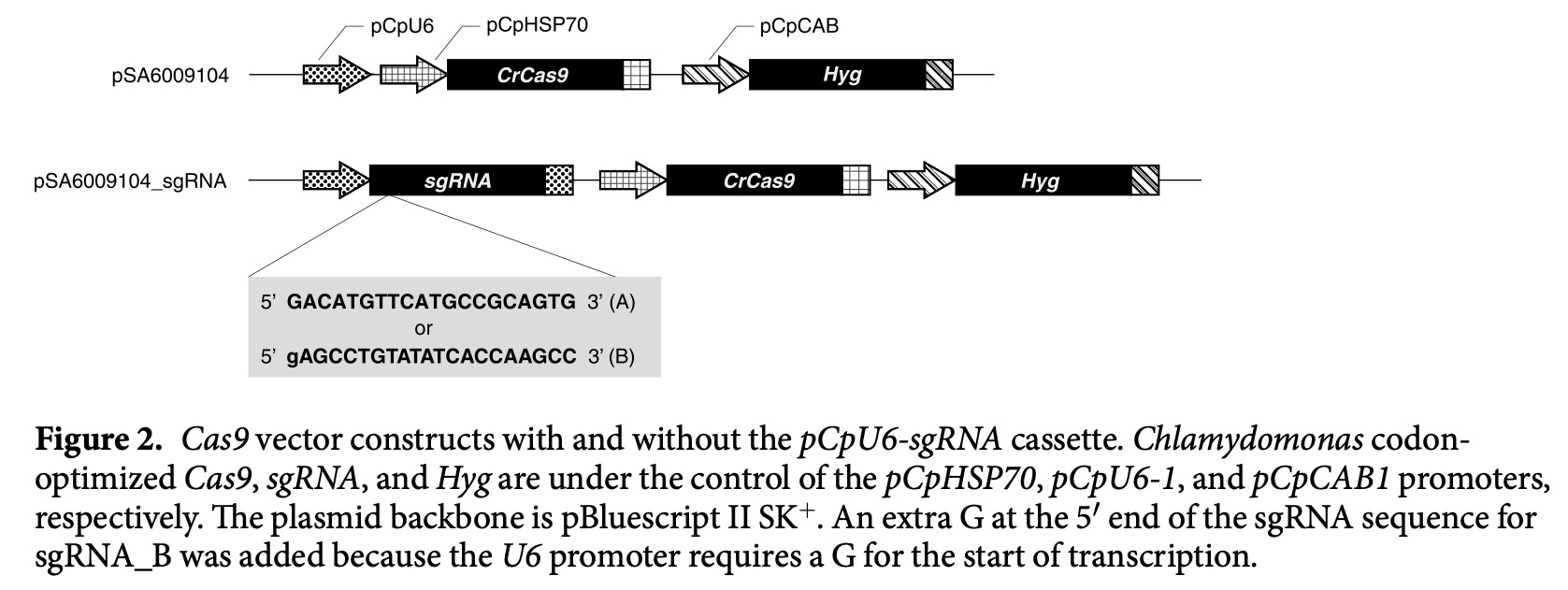
The mt+ and mt– cells recognize each other by chemical communication through mating-type-specific sex pheromones: protoplast-release-inducing protein (PR-IP) Inducer and PR-IP (Fig. 2, Akatsuka et al. 2003, 2006; Tsuchikane et al. 2003; Sekimoto et al. 2012). PR-IP Inducer is a glycoprotein (Nojiri et al. 1995) that is released from the mt– cells and induces both SCD (Tsuchikane et al. 2005) and PR-IP production (Sekimoto et al. 1994; Sekimoto 2002) in mt+ cells. PR-IP is also a glycoprotein that is released from the mt+ cells and induces both the SCD (Akatsuka et al. 2006) and the release of gametic protoplasts from mt– cells (Sekimoto et al. 1990). The cDNAs encoding the respective pheromones have been cloned (Sekimoto et al. 1994, 1998).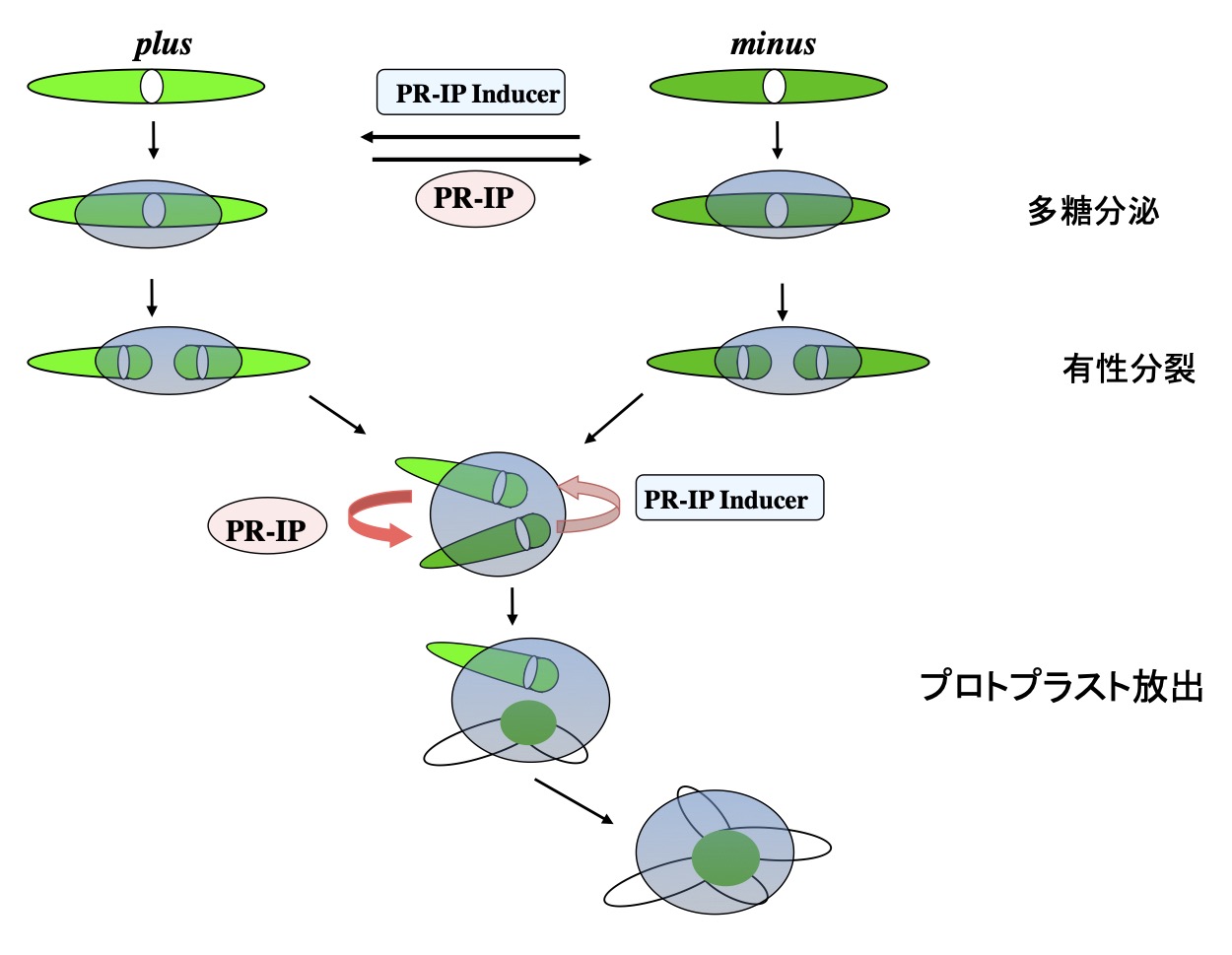
2)Microarray analyses to identify mating-related genes
To identify mating-related genes, we had constructed a cDNA microarray. As a result, some of sex-specific, pheromone-inducible, or conjugation-related genes had been detected (Sekimoto et al. 2003, 2006).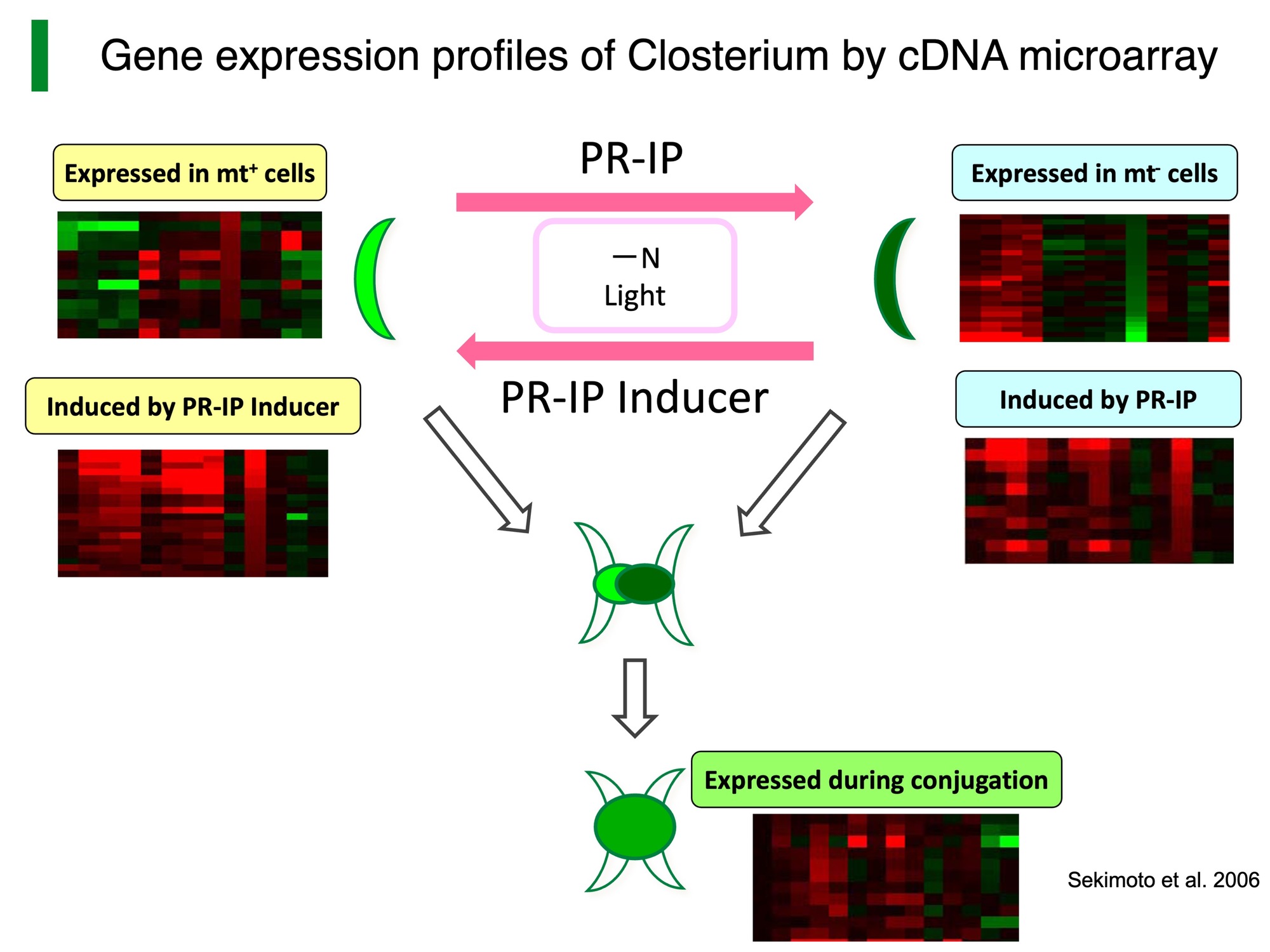
3)Development of stable transformation system for the Closterium peracerosum-strigosum-littorale complex
We previously generated the first stable genetic transformants of the unicellular zygnematophycean alga Closterium peracerosum-strigosum-littorale (C. psl.) complex using particle bombardment (Abe et al., 2011). However, there are some difficulties in the genetic transformation of C. psl. complex using the method. First, the transformants are often contaminated by bacteria and/or fungi, making it necessary to wash and isolate single cells using glass capillaries under a microscope. This operation sometimes results in additional contamination of (or injury to) the transformants. Second, the efficiency of transformation is not very high; the average transformation efficiency following selection in 0.3 µg mL-1 phleomycin was 5.5 x 10-6 cells or 0.03 transformants per µg DNA(Abe et al., 2011). Therefore, many cells must be prepared in order to establish a sufficient number of transformants (Abe et al., 2016), which requires a large amount of incubator space. We have tried to establish other transformation methods, such as electroporation using protoplasts. However, normal lunate cells could not be regenerated from the protoplasts irrespective of the electric stimulus.
To overcome these difficulties, we developed a transformation system using a square electric pulse–generating electroporator and succeeded in establishing a highly efficient transformation technique for wild C. psl. complex without the need for cell-wall removal. To date, this technique is the only reliable method for the transformation of charophycean algae.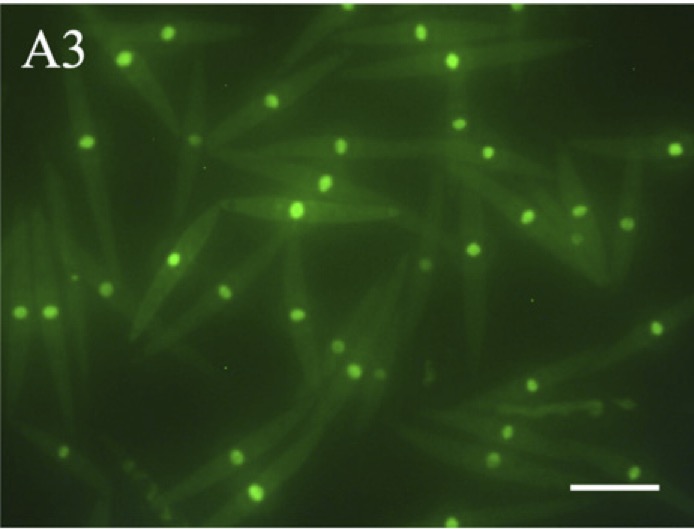
4)Physiological function of CpRLK1
One of the genes identified by the microarray analysis encodes a receptor-like protein kinase (RLK) and is named CpRLK1. The gene is expressed specifically in mt+ cells. The expression is elevated during sexual reproduction and treatment of mt+ cells with the PR-IP Inducer also promotes its expression, indicating that the CpRLK1 protein probably functions during sexual reproduction (Sekimoto et al., 2006). Mt+ cells with knockdown of CpRLK1 showed reduced competence for sexual reproduction and formed an abnormally enlarged conjugation papilla after pairing with mt- cells. The knockdown cells were unable to release a naked gamete, which is indispensable for zygote formation. We suggest that the CpRLK1 protein is an ancient cell wall sensor that now functions to regulate osmotic pressure in the cell to allow proper gamete release.
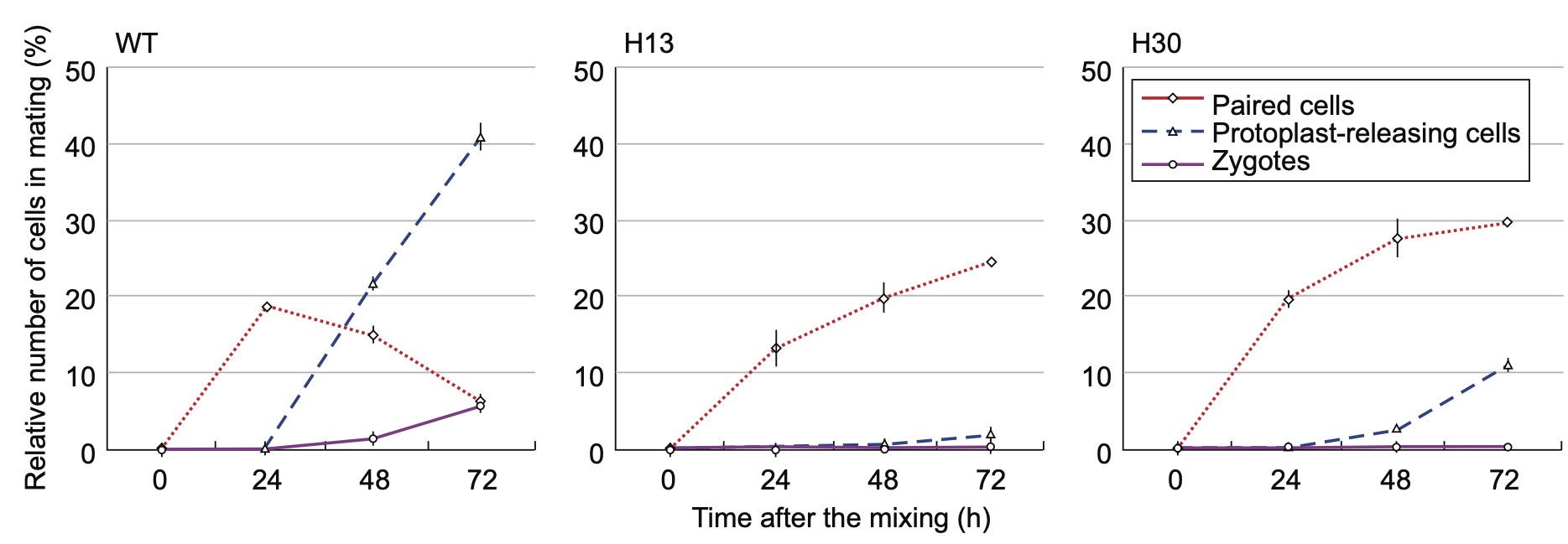
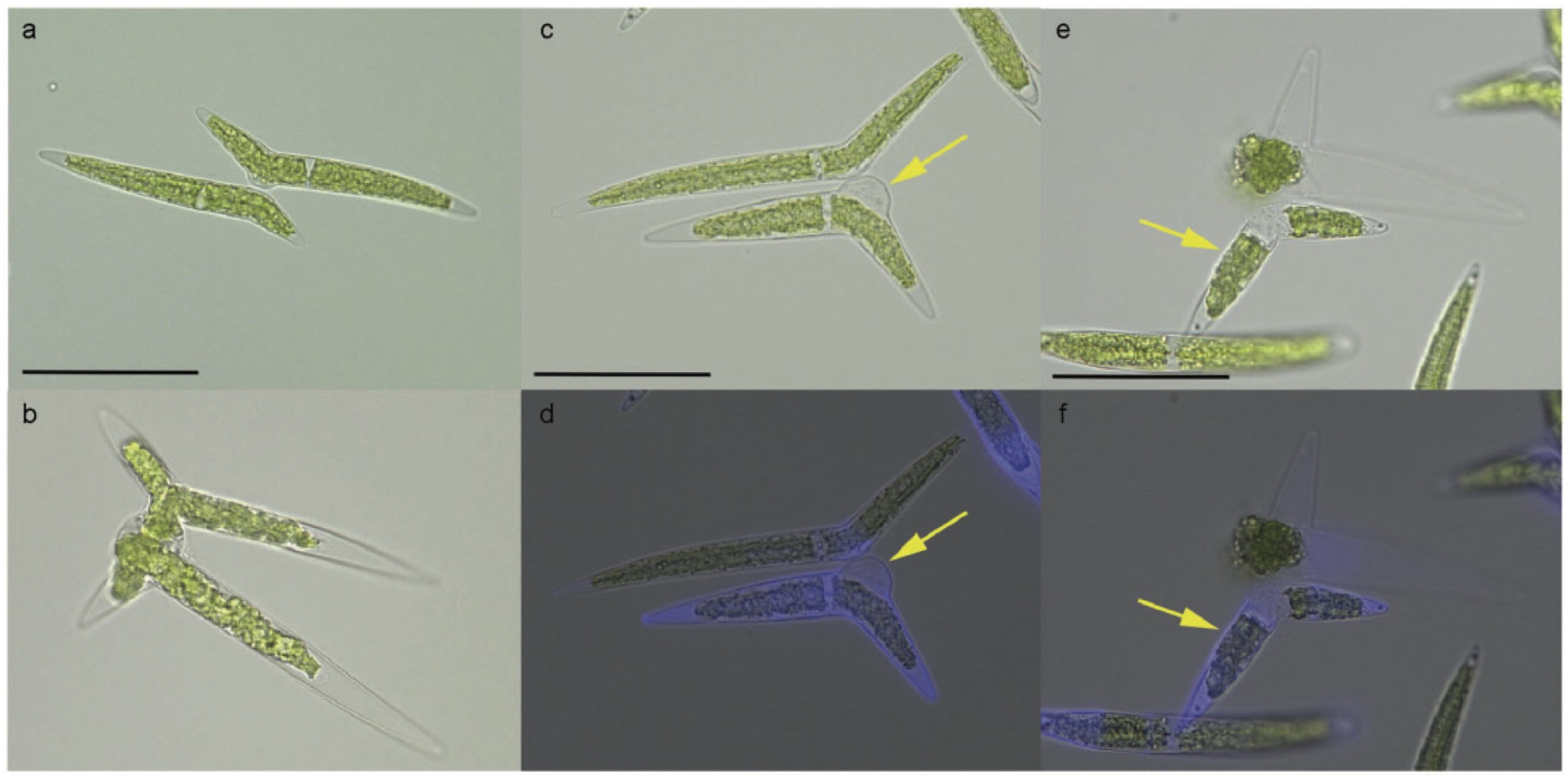
5)Establishment of CRISPR/Cas9-based gene knockout system and evaluation of CpRLP1
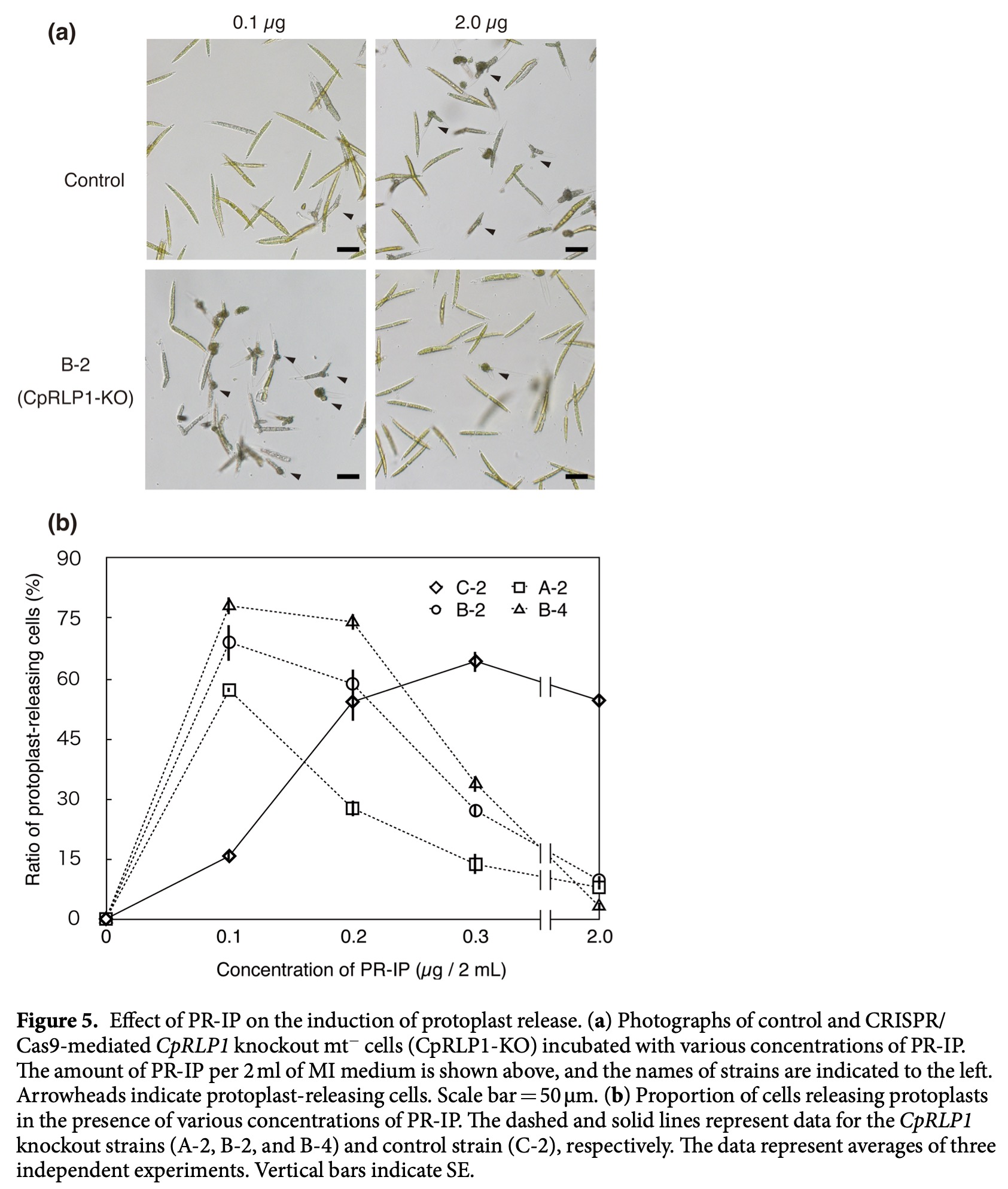
We examined the mechanism of action of CpRLP1 (receptor-like protein 1), which was previously identified in a cDNA microarray analysis as one of the PR-IP-inducible genes. Using CRISPR/Cas9 technology, we generated CpRLP1knockout mutants in mt– cells of the C. psl. complex. When the knockout mt– cells were mixed with wild-type mt+ cells, conjugation was severely reduced. Many cells released protoplasts without pairing, suggesting a loss of synchronization between the two mating partners. Furthermore, the knockout mutants were hypersensitive to PR-IP. We conclude that CpRLP1 is a negative regulator of PR-IP that regulates the timing of protoplast release in conjugating C. psl. cells. As the first report of successful gene knockout in the class Charophyceae, this study provides a basis for research aimed at understanding the ancestral roles of genes that are indispensable for the development of land plants.
6)Whole genome sequencing reveals that a divergent RWP-RK transcription factor determines mating type in heterothallic Closterium
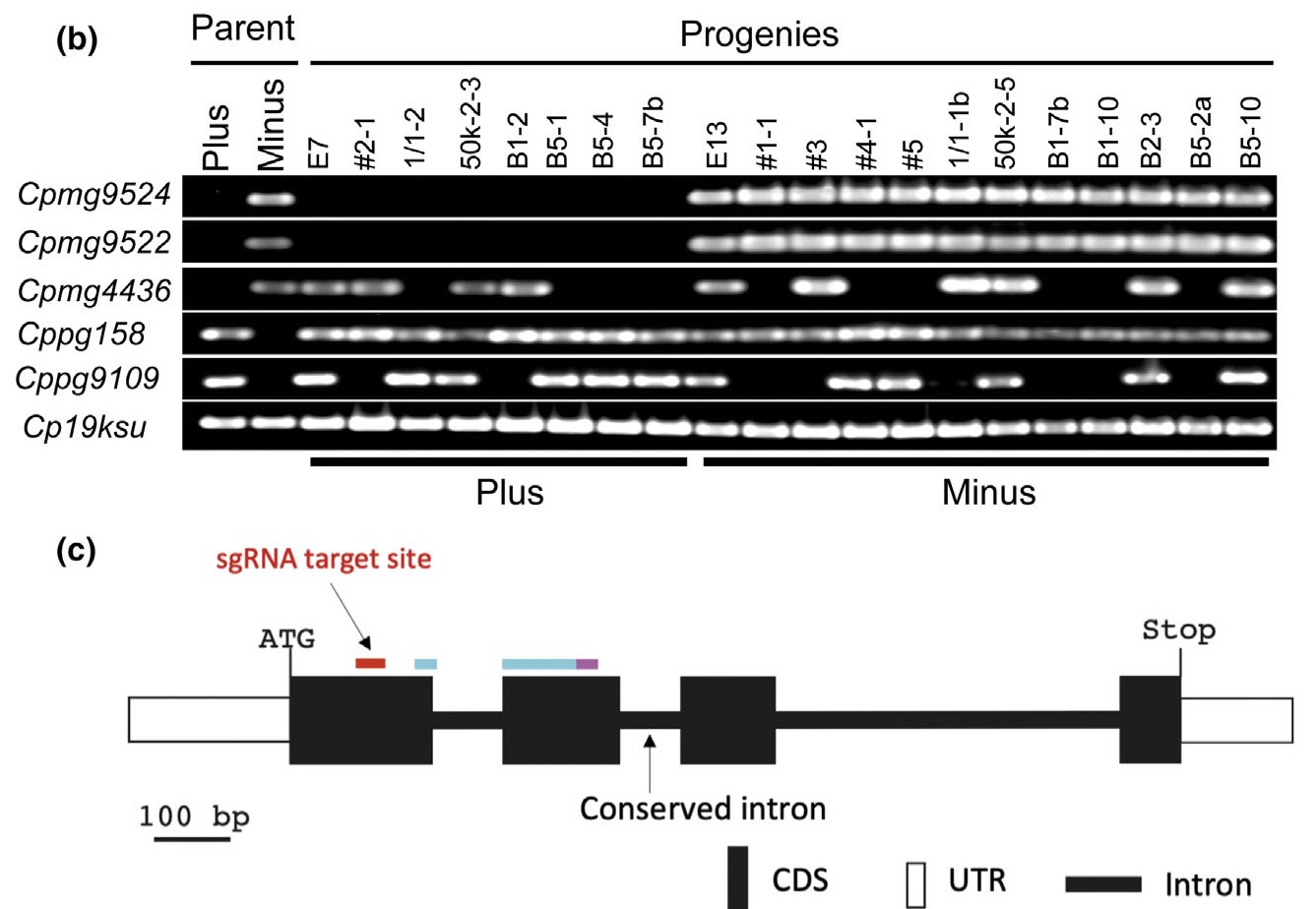
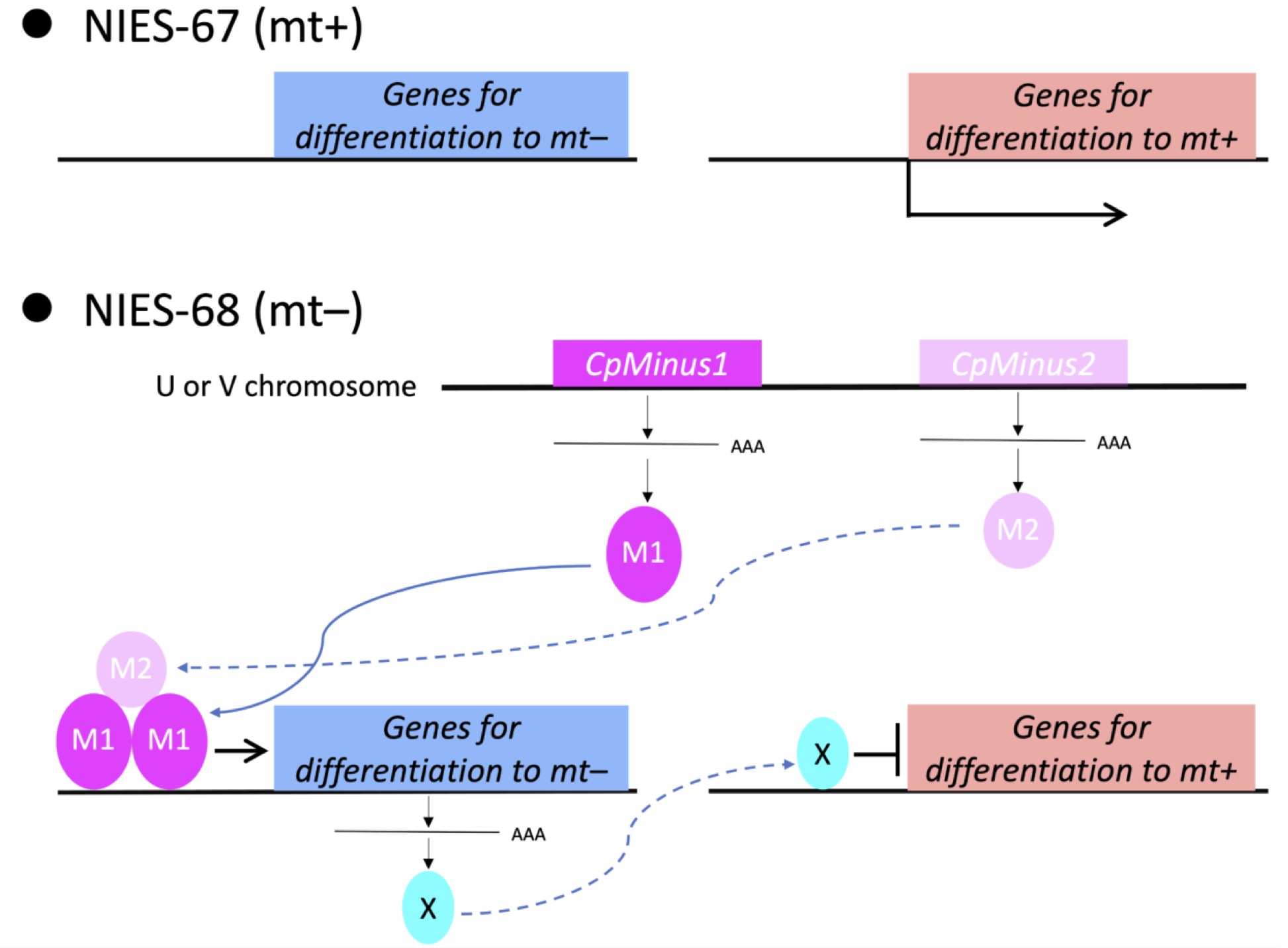
The Closterium has an isogamous mating system. We sequenced genomes representing the two MTs, mt+ and mt–, in Closterium and identified CpMinus1, a gene linked to the mt– phenotype. CpMinus1 encodes a divergent RWP-RK domain–containing (RKD)-like transcription factor and is specifically expressed during gamete differentiation. Introduction of CpMinus1 into an mt+ strain was sufficient to convert it to a phenotypically mt– strain, while CpMinus1-knockout mt– strains were phenotypically mt+. We propose that CpMinus1 is the major MT determinant that acts by evoking the mt– phenotype and suppressing the mt+ phenotype in heterothallic Closterium. CpMinus1 likely evolved independently in the Zygnematophyceae lineage, which lost an egg-sperm anisogamous mating system. mt–-specific regions possibly constitute an MT locus flanked by common sequences that undergo some recombination.